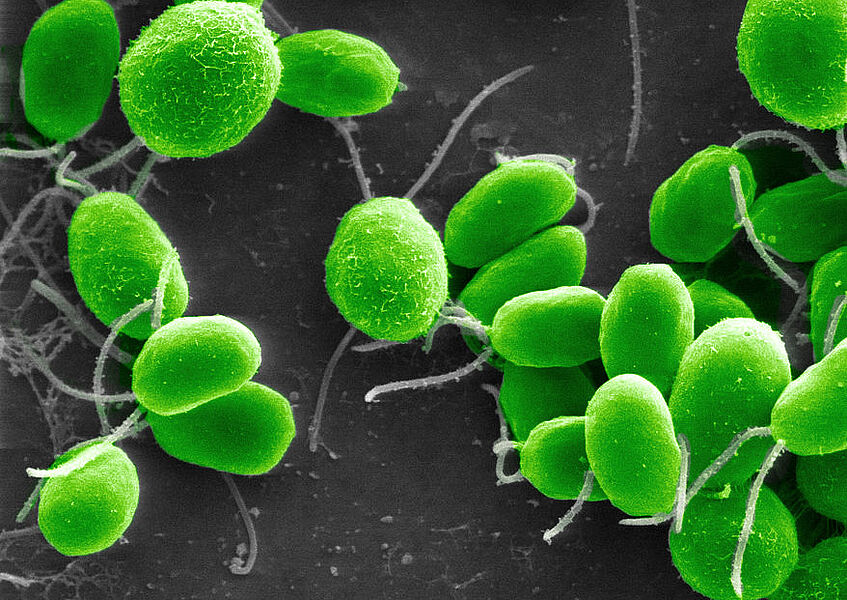
Chlamydomonas reinhardtii - model system for alternative energy resources
Plant biology and biotechnology has become dramatically important in dealing with challenges of
- world population feeding,
- global climate change and
- limited energy resources with fossil fuels.
One key issue is the development of CO2-neutral plant resources for biomass and biofuels: a transition from first generation plants like sugar cane, maize and other important nutritional crops to second and third generation energy systems such as algae for biomass and feed, hydrogen and lipid production.
The algae Chlamydomonas reinhardtii is an early and prominent model system for third generation energy resources which prompted us to do extended research on physiology and signaling during the life cycle of this fascinating unicellular green organism.
Introducion to the topic of Green systems biology
Green systems biology - From single genomes, proteomes and metabolomes to ecosystems research and biotechnology. Weckwerth W.J Proteomics. 2011 Dec 10;75(1):284-305. doi: 10.1016/j.jprot.2011.07.010. Epub 2011 Jul 23. Review.
Research on Chlamydomonas reinhardtii
- Biodiesel and poly-unsaturated fatty acids production from algae and crop plants - a rapid and comprehensive workflow for lipid analysis. Furuhashi T, Nakamura T, Fragner L, Roustan V, Schön V, Weckwerth W. Biotechnol J. 2016 Oct;11(10):1262-1267. doi: 10.1002/biot.201400197. Epub 2016 Aug 11.
- ProMEX: a mass spectral reference database for proteins and protein phosphorylation sites. Hummel J, Niemann M, Wienkoop S, Schulze W, Steinhauser D, Selbig J, Walther D, Weckwerth W. BMC Bioinformatics. 2007 Jun 23;8:216.
- An automated GCxGC-TOF-MS protocol for batch-wise extraction and alignment of mass isotopomer matrixes from differential 13C-labelling experiments: a case study for photoautotrophic-mixotrophic grown Chlamydomonas reinhardtii cells. Kempa S, Hummel J, Schwemmer T, Pietzke M, Strehmel N, Wienkoop S, Kopka J, Weckwerth W. J Basic Microbiol. 2009 Feb;49(1):82-91. doi: 10.1002/jobm.200800337.
- Plant protein phosphorylation monitored by capillary liquid chromatography--element mass spectrometry. Krüger R, Wolschin F, Weckwerth W, Bettmer J, Lehmann WD. Biochem Biophys Res Commun. 2007 Mar 30;355(1):89-96. Epub 2007 Jan 31.
- Metabolomics- and proteomics-assisted genome annotation and analysis of the draft metabolic network of Chlamydomonas reinhardtii. May P, Wienkoop S, Kempa S, Usadel B, Christian N, Rupprecht J, Weiss J, Recuenco-Munoz L, Ebenhöh O, Weckwerth W, Walther D. Genetics. 2008 May;179(1):157-66. doi: 10.1534/genetics.108.088336.
- Targeted quantitative analysis of a diurnal RuBisCO subunit expression and translation profile in Chlamydomonas reinhardtii introducing a novel Mass Western approach. Recuenco-Muñoz L, Offre P, Valledor L, Lyon D, Weckwerth W, Wienkoop S. J Proteomics. 2015 Jan 15;113:143-53. doi: 10.1016/j.jprot.2014.09.026. Epub 2014 Oct 7.
- Quantitative in vivo phosphoproteomics reveals reversible signaling processes during nitrogen starvation and recovery in the biofuel model organism Chlamydomonas reinhardtii. Roustan V, Bakhtiari S, Roustan PJ, Weckwerth W. Biotechnol Biofuels. 2017 Nov 28;10:280. doi: 10.1186/s13068-017-0949-z. eCollection 2017.
- A universal protocol for the combined isolation of metabolites, DNA, long RNAs, small RNAs, and proteins from plants and microorganisms. Valledor L, Escandón M, Meijón M, Nukarinen E, Cañal MJ, Weckwerth W. Plant J. 2014 Jul;79(1):173-80. doi: 10.1111/tpj.12546. Epub 2014 Jun 17.
- System-level network analysis of nitrogen starvation and recovery in Chlamydomonas reinhardtii reveals potential new targets for increased lipid accumulation. Valledor L, Furuhashi T, Recuenco-Muñoz L, Wienkoop S, Weckwerth W. Biotechnol Biofuels. 2014 Dec 24;7:171. doi: 10.1186/s13068-014-0171-1. eCollection 2014.
- An improved detergent-compatible gel-fractionation LC-LTQ-Orbitrap-MS workflow for plant and microbial proteomics. Valledor L, Weckwerth W. Methods Mol Biol. 2014;1072:347-58. doi: 10.1007/978-1-62703-631-3_25.
- Systemic cold stress adaptation of Chlamydomonas reinhardtii. Valledor L, Furuhashi T, Hanak AM, Weckwerth W. Mol Cell Proteomics. 2013 Aug;12(8):2032-47. doi: 10.1074/mcp.M112.026765. Epub 2013 Apr 5.
- The different proteomes of Chlamydomonas reinhardtii. Valledor L, Recuenco-Munoz L, Egelhofer V, Wienkoop S, Weckwerth W. J Proteomics. 2012 Oct 22;75(18):5883-7. doi: 10.1016/j.jprot.2012.07.045. Epub 2012 Aug 7.
- ProMEX - a mass spectral reference database for plant proteomics. Wienkoop S, Staudinger C, Hoehenwarter W, Weckwerth W, Egelhofer V. Front Plant Sci. 2012 Jun 6;3:125. doi: 10.3389/fpls.2012.00125. eCollection 2012.
- Targeted proteomics for Chlamydomonas reinhardtii combined with rapid subcellular protein fractionation, metabolomics and metabolic flux analyses. Wienkoop S, Weiss J, May P, Kempa S, Irgang S, Recuenco-Munoz L, Pietzke M, Schwemmer T, Rupprecht J, Egelhofer V, Weckwerth W. Mol Biosyst. 2010 Jun;6(6):1018-31. doi: 10.1039/b920913a. Epub 2010 Mar 31.
- Combining metal oxide affinity chromatography (MOAC) and selective mass spectrometry for robust identification of in vivo protein phosphorylation sites. Wolschin F, Weckwerth W. Plant Methods. 2005 Nov 1;1(1):9.
